✨ Project Summary
My research spans four interconnected domains: Genomics, Computational Drug Discovery, Immunoinformatics, and Wet-Lab Experimental Biology. I integrate molecular techniques with advanced computational workflows to investigate disease-associated variants, design therapeutic candidates, and develop immunoinformatics-driven vaccines.
12+ research projects • 5+ major disease targets • 4 interdisciplinary domains
🧬 1. Genomics & Variant Analysis
MMP3 & MMP9 Variant Profiling in Periodontitis and Diabetes
Identifying disease-associated MMP variants using wet-lab and computational workflows.
Highlights:
Highlights:
- PCR amplification, Sanger sequencing
- NGS-based variant calling
- Functional annotation (GATK, VEP, ClinVar)
- Novel variants identified across Bangladeshi population
Whole Exome Sequencing: Variant Discovery Pipeline
High-throughput WES analysis for disease-associated SNPs and INDELs.
Highlights:
Highlights:
- QC: FASTQC, Trimmomatic
- Alignment: Hisat2 / BWA
- Variant calling: GATK
- Downstream analysis: VCFtools, SnpEff
Breast Cancer GWAS-Based Variant Annotation (In preparation)
Screening population-level SNPs to identify risk-associated loci.
Highlights:
Highlights:
- GWAS dataset curation
- Manhattan and QQ plot analysis
- Functional annotation & pathogenicity scoring

💻 2. Computational Drug Discovery
Structure-Based Drug Design Against Chikungunya RdRp
First-author Q1 publication on lead compound identification targeting CHIKV RdRp.
Highlights:
- Protein preparation & active-site mapping
- Virtual screening & docking (AutoDock, PyRx)
- MD simulation (GROMACS)
- ADMET & DFT validation
P53 Nanophytocompound Screening & MD Validation
Investigating natural compounds targeting P53 for therapeutic potential.
Highlights:
Highlights:
- Ligand screening & in silico toxicity prediction
- 100 ns MD simulation for stability analysis
- RMSD/RMSF free energy profiling
- Animal model bioactivity testing (wet-lab validation)
Lead Optimization for Viral Targets
Computational identification of small molecules targeting viral membrane and polymerase proteins.
Highlights:
Highlights:
- Molecular docking
- Energy minimization
- Hotspot analysis
- Drug-likeness scoring

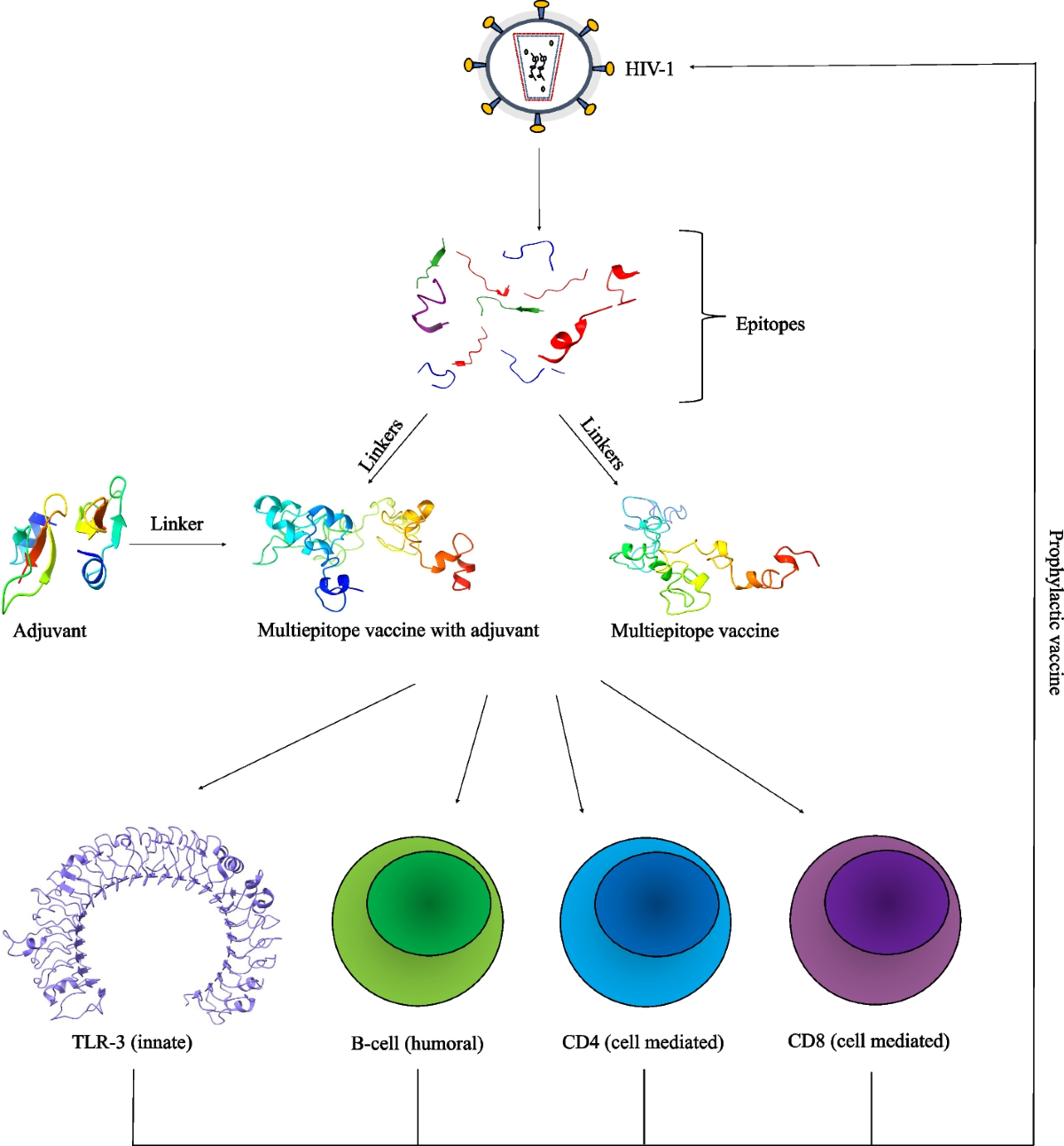
💉 3. Immunoinformatics & Vaccine Design
Mpox Multi-Epitope Vaccine Design (Q1 publication)
A complete immunoinformatics pipeline to generate epitope-based vaccine candidates.
Highlights:
Highlights:
- CTL, HTL, B-cell epitope screening
- Molecular docking with immune receptors
- Allergenicity & antigenicity profiling
- In silico cloning
Reverse Vaccinology for C. trachomatis (Submitted)
Computational antigen discovery and epitope prioritization.
Highlights:
Highlights:
- Proteome mining
- Subcellular localization prediction
- B/T-cell epitope mapping
- Population coverage analysis
Vaccine Candidate Selection for S. pyogenes
Identifying conserved epitopes for rational vaccine design.
Highlights:
Highlights:
- Antigenic region mapping
- Conservancy analysis
- Adjuvant design
- 3D modeling & immunological simulation
🧪 4. Wet Lab Research & Internships
Sanger Sequencing & NGS Training — NIB
Hands-on experience with Sanger workflow and next-generation sequencing concepts.
Highlights:
Highlights:
- DNA extraction & quantification
- PCR optimization
- Gel electrophoresis
- Sequence analysis
Nanophytocompound Extraction & Animal Testing
Experimental validation of two lead compounds with high binding affinity to P53.
Highlights:
Highlights:
- Plant extract preparation
- Column chromatography
- Animal model toxicity & bioactivity testing
- Correlation with computational predictions
Undergraduate Research: CHIKV RdRp Targeting
Complete bioinformatics pipeline for antiviral drug discovery.
Highlights:
Highlights:
- Research design
- Protein modelling
- Docking & MD simulation
- HOMO–LUMO, DFT studies
